2.2 Binding basics
Consider this code:
x <- c(1, 2, 3)It’s easy to read it as: “create an object named ‘x’, containing the values 1, 2, and 3”. Unfortunately, that’s a simplification that will lead to inaccurate predictions about what R is actually doing behind the scenes. It’s more accurate to say that this code is doing two things:
- It’s creating an object, a vector of values,
c(1, 2, 3). - And it’s binding that object to a name,
x.
In other words, the object, or value, doesn’t have a name; it’s actually the name that has a value.
To further clarify this distinction, I’ll draw diagrams like this:
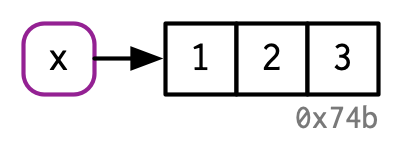
The name, x, is drawn with a rounded rectangle. It has an arrow that points to (or binds or references) the value, the vector c(1, 2, 3). The arrow points in opposite direction to the assignment arrow: <- creates a binding from the name on the left-hand side to the object on the right-hand side.
Thus, you can think of a name as a reference to a value. For example, if you run this code, you don’t get another copy of the value c(1, 2, 3), you get another binding to the existing object:
y <- x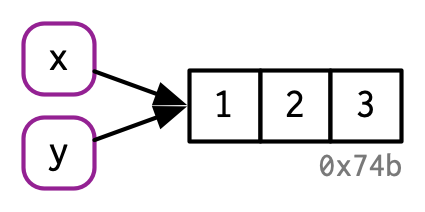
You might have noticed that the value c(1, 2, 3) has a label: 0x74b. While the vector doesn’t have a name, I’ll occasionally need to refer to an object independent of its bindings. To make that possible, I’ll label values with a unique identifier. These identifiers have a special form that looks like the object’s memory “address”, i.e. the location in memory where the object is stored. But because the actual memory addresses changes every time the code is run, we use these identifiers instead.
You can access an object’s identifier with lobstr::obj_addr(). Doing so allows you to see that both x and y point to the same identifier:
obj_addr(x)
#> [1] "0x55d537cd7888"
obj_addr(y)
#> [1] "0x55d537cd7888"These identifiers are long, and change every time you restart R.
It can take some time to get your head around the distinction between names and values, but understanding this is really helpful in functional programming where functions can have different names in different contexts.
2.2.1 Non-syntactic names
R has strict rules about what constitutes a valid name. A syntactic name must consist of letters2, digits, . and _ but can’t begin with _ or a digit. Additionally, you can’t use any of the reserved words like TRUE, NULL, if, and function (see the complete list in ?Reserved). A name that doesn’t follow these rules is a non-syntactic name; if you try to use them, you’ll get an error:
_abc <- 1
#> Error: unexpected input in "_"
if <- 10
#> Error: unexpected assignment in "if <-"It’s possible to override these rules and use any name, i.e., any sequence of characters, by surrounding it with backticks:
`_abc` <- 1
`_abc`
#> [1] 1
`if` <- 10
`if`
#> [1] 10While it’s unlikely you’d deliberately create such crazy names, you need to understand how these crazy names work because you’ll come across them, most commonly when you load data that has been created outside of R.
2.2.2 Exercises
Explain the relationship between
a,b,canddin the following code:a <- 1:10 b <- a c <- b d <- 1:10The following code accesses the mean function in multiple ways. Do they all point to the same underlying function object? Verify this with
lobstr::obj_addr().mean base::mean get("mean") evalq(mean) match.fun("mean")By default, base R data import functions, like
read.csv(), will automatically convert non-syntactic names to syntactic ones. Why might this be problematic? What option allows you to suppress this behaviour?What rules does
make.names()use to convert non-syntactic names into syntactic ones?I slightly simplified the rules that govern syntactic names. Why is
.123e1not a syntactic name? Read?make.namesfor the full details.
Surprisingly, precisely what constitutes a letter is determined by your current locale. That means that the syntax of R code can actually differ from computer to computer, and that it’s possible for a file that works on one computer to not even parse on another! Avoid this problem by sticking to ASCII characters (i.e. A-Z) as much as possible.↩︎